Very happy to share our recent study on the application of an integrated multi-level computational approach to identify new promising molecules to selective inhibit biofilm formation by Pseudomonas aeruginosa, published in RSC’s journal Molecular Systems Design & Engineering (IF 4.92). Great work by Tatiana F. Vieira, as part of her PhD work, in collaboration with Prof. Manuel Simões (FEUP).
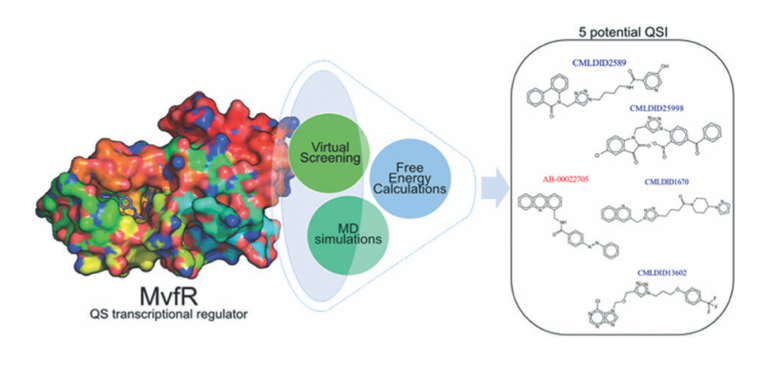
Targeting Pseudomonas aeruginosa MvfR in the battle against biofilm formation: a multi-level computational approach
Tatiana F. Vieira, Rita P. Magalhães, Nuno M. F. S. A. Cerqueira, Manuel Simões and Sérgio F. Sousa
Molecular Systems Design & Engineering | DOI: 10.1039/D2ME00088A
Abstract:
Resistance to antibiotics is an increasing threat in global health and strategies to develop or find new drugs are of the utmost importance. Pseudomonas aeruginosa is a Gram-negative pathogen that shows resistance to therapeutics and that leads to high morbidity and mortality rates. Through a multiple virulence factor regulator (MvfR or PqsR) it can regulate the transcription of several virulence genes responsible for the formation of biofilms and antibiotic-tolerant/persister (AT/P) cells. In this work, four large databases of compounds – Mu.Ta.Lig. Virtual Chemotheca, Chimiothèque Nationale, Comprehensive Marine Natural Products Database (CMNPD) and the natural compounds database of InterbioScreen (IBS) (representing a total of 217 939 compounds), were screened against MvfR and 20 compounds were identified as potential inhibitors for further experimental validation. Molecular dynamics simulations and free energy calculations were performed to validate the docking predictions and estimate binding free energies. Of the 20 compounds, 13 have shown better total free energies of association than the reference molecule (M64, an antagonist). After the prediction of ADMET and pharmacokinetic properties, a final list composed of seven more promising quorum sensing inhibitors was selected. A detailed characterization of the binding modes of each of these molecules was also presented providing clues for future developments.

