Please check our new research article published in Molecular Systems Design & Engineering (IF: 4.94), a collaboration with Professor Lígia Rodrigues’ group (CEB, University of Minho). In this paper we applied docking, molecular dynamics simulations, and binding free energy calculations with MM/GBSA for the identification of novel aptamers targeting prostate cancer cells overexpressing cathepsin B. These results were further validated by cell binding assays, corroborating the in silico predictions.
Identification of novel aptamers targeting cathepsin B-overexpressing prostate cancer cells
Ana Cláudia Pereira,‡ André F. Pina,‡ Diana Sousa,‡ Débora Ferreira, Cátia Santos-Pereira, Joana L. Rodrigues, Luís D. R. Melo, Goreti Sales, Sérgio F. Sousa and Lígia R. Rodrigues
MSDE (2022) | DOI: 10.1039/D2ME00022A
‡ The authors contributed equally to the work.
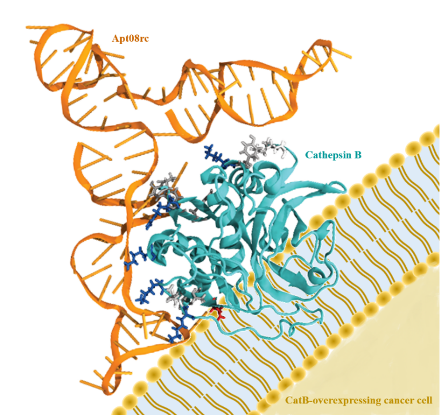
Abstract
Increased levels of cathepsin B (CatB), a cysteine protease, have been associated with different types of tumors, including prostate cancer. Hence, the identification of novel targeting ligands homing CatB may be a promising approach for the development of CatB-targeted therapies. In this work, a methodology called systematic evolution of ligands by EXponential enrichment (SELEX) was used to generate and isolate new aptamers targeting CatB. After 8 rounds of selection, the pool was sequenced, and the 10 most prevalent sequences, along with their corresponding reverse complementary sequences, were further analyzed. In order to assess the binding affinity of the aptamers towards the CatB, bioinformatics tools were used. After generating the 3D structures for each aptamer, the HADDOCK software was used for the docking studies between them and CatB. Molecular dynamics simulations and free binding energy calculations by molecular mechanics-generalized born surface area (MM-GBSA) allowed selection of a new aptamer with potential to target CatB. Cell binding assays against the PC-3 prostate cancer cell line confirmed the in silico predictions, further validating the newly discovered aptamer as a promising targeting ligand for CatB-overexpressing cancer cells.
